Personalized Science
We help scientists analyze the most relevant gene expression data for their questions
Our mission is to enable scientists to treat and cure disease
We build software to enable biologists to use their knowledge and expertise to discover and validate targets via the analysis and exploration of highly relevant gene expression datasets. Founded by Brian Fox LinkedIn
Personalized Science: Every scientific team has a different focus. We are not aiming to make every piece of expression data available, but rather the key, high-quality datasets in your area of research. It is up to you how personalize your server instance with specific datasets and analyses.
Features
Our gene expression app is personalized for you.
Intuitive
Designed with the inquisitive biologist in mind. We focus on gene expression (bulk and single cell) because of the high re-use value and it is fundamentally approachable compared to other omics data types.
Browser Based
Use any laptop to browse the data. No need for users to install anything.
Cloud or on-prem
Use a shared cloud instance or a custom install on servers you control, within your firewall. The server app is built using django and django rest framework and can be installed via docker or we can provide a pre-configured EBS volume.
R scripts
Most of the plots and many of the analyses use R and various open-source and community built packages: no secret methods. Although we use ggplot, we do NOT use R-Shiny for the user interface. The R API allows you to perform your own analyses in R and easily grab any data from the server database.
Data Library
We have a library of over 100 public datasets and over 1M cells that you can select for your instance of the app. The app has curation tools for adding new GEO datasets very quickly and easily.
Development
Your needs can be prioritized in upcoming development sprints of the app. We love working closely with fellow scientists and solving their problems.
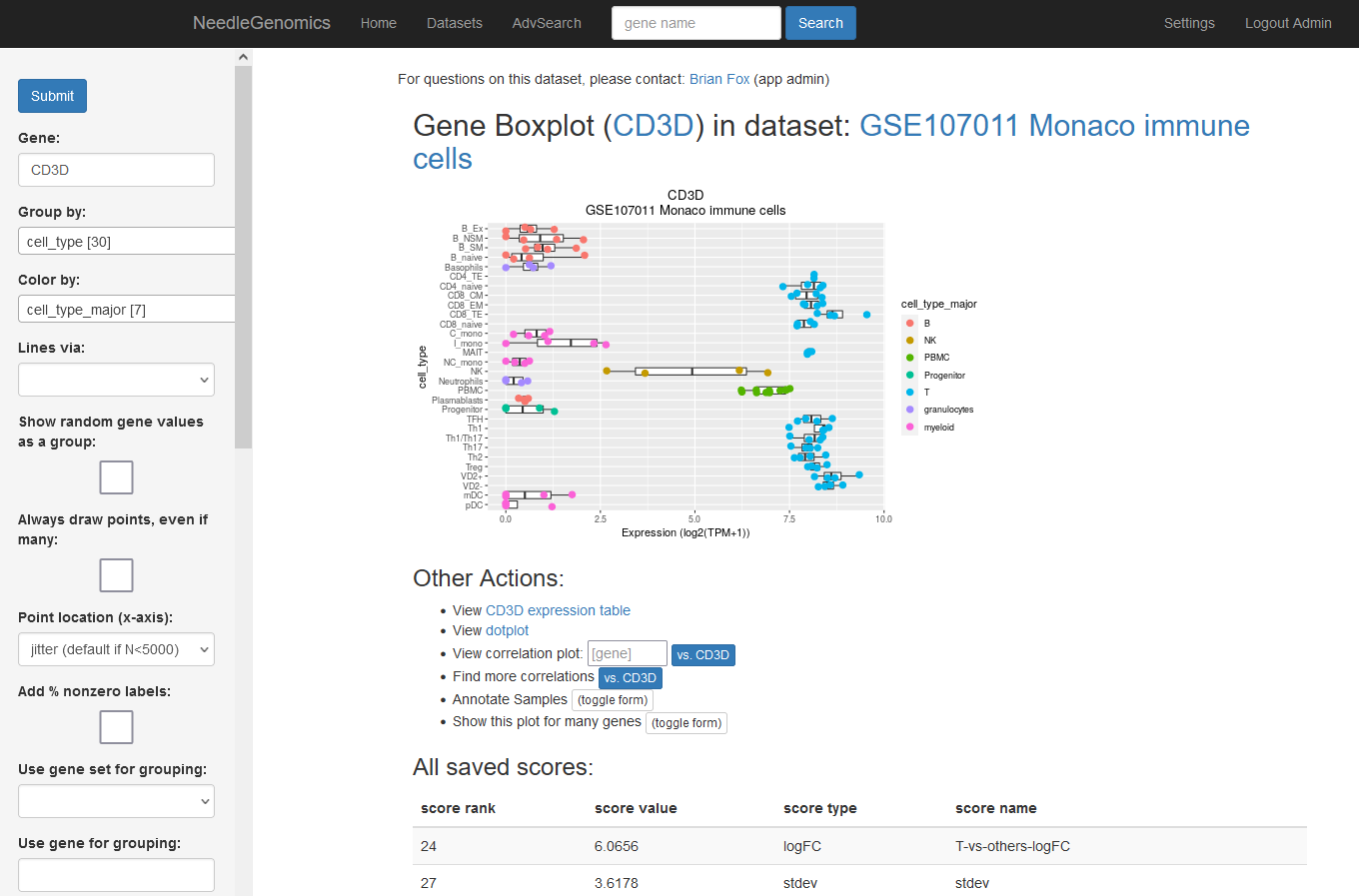
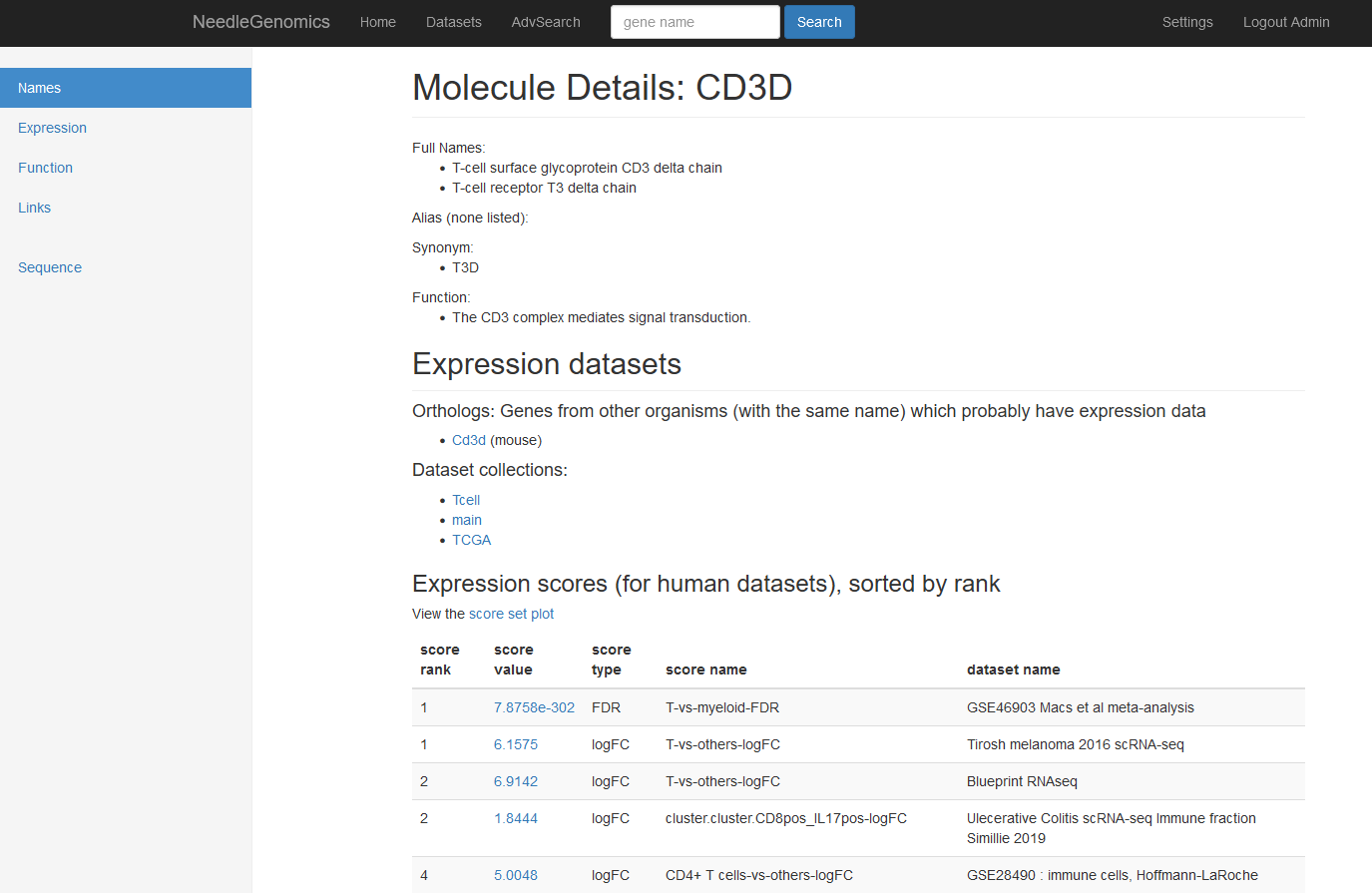
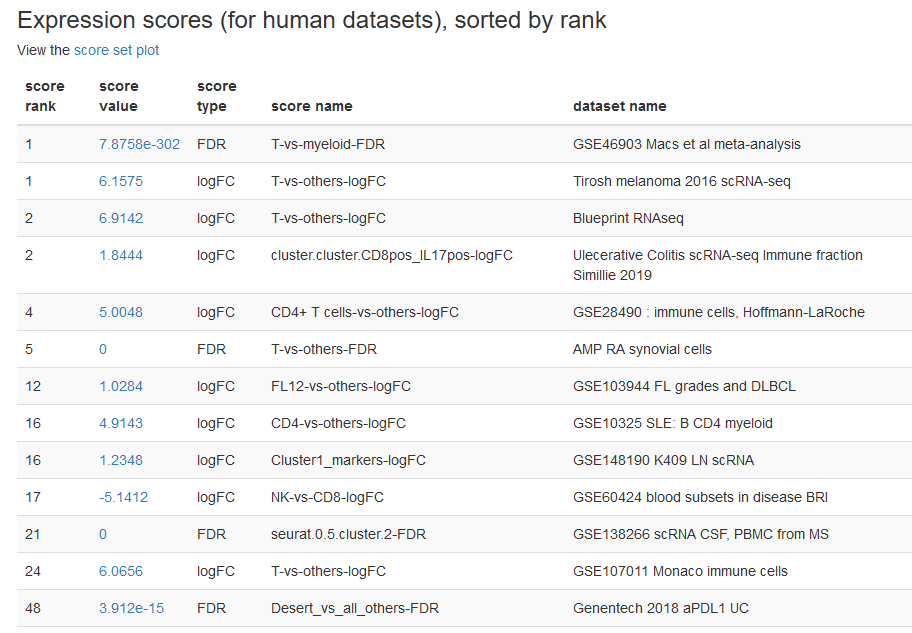
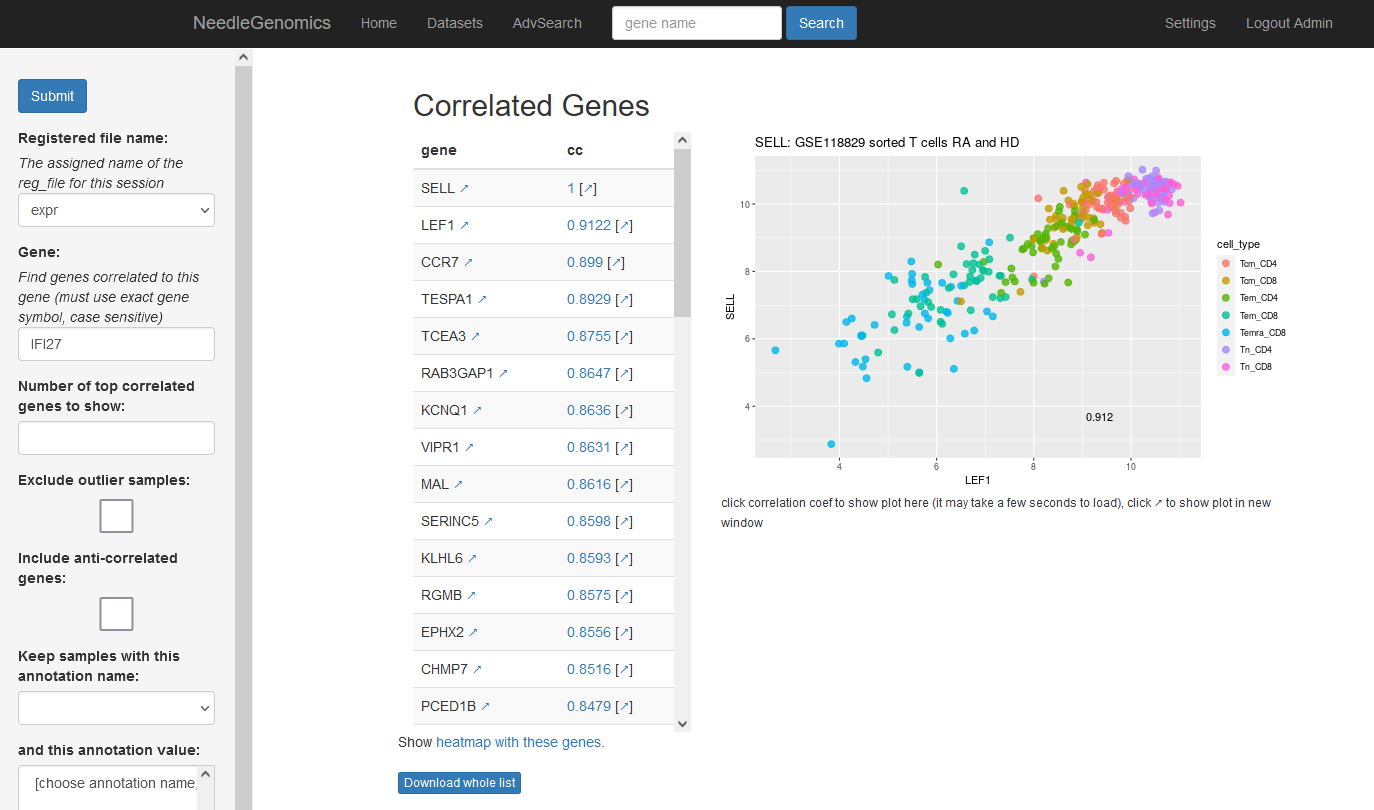
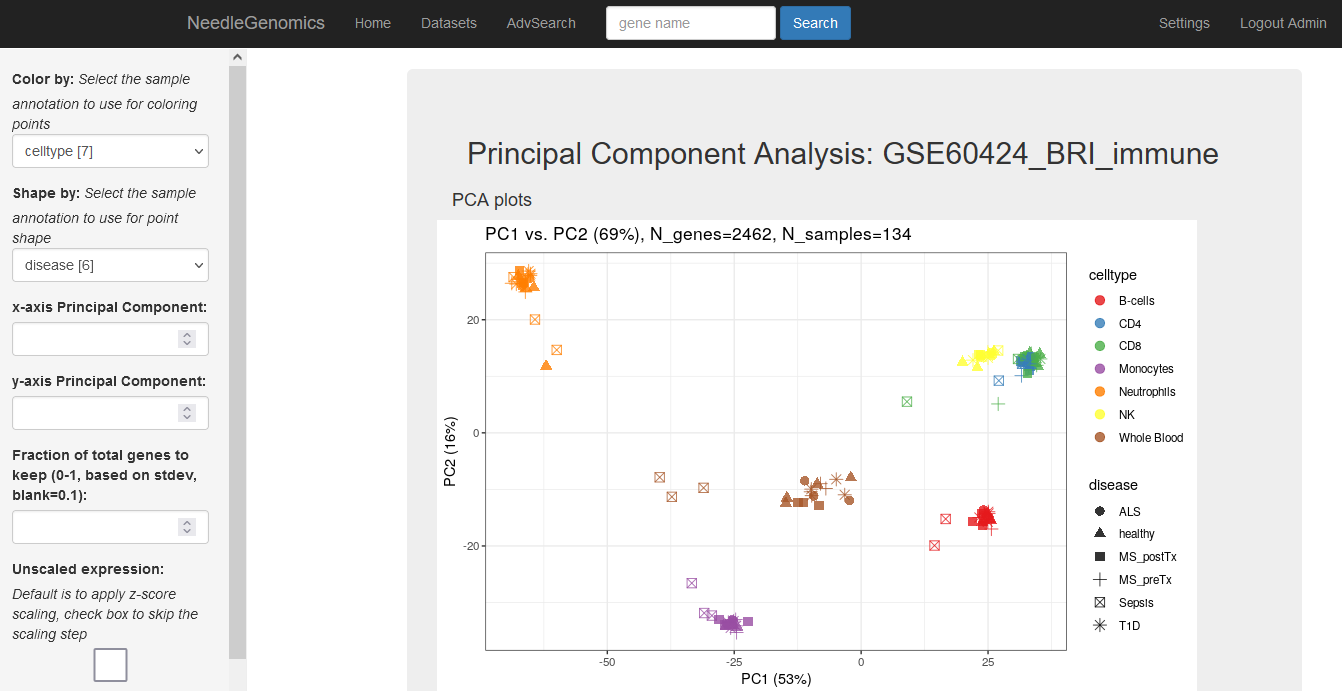
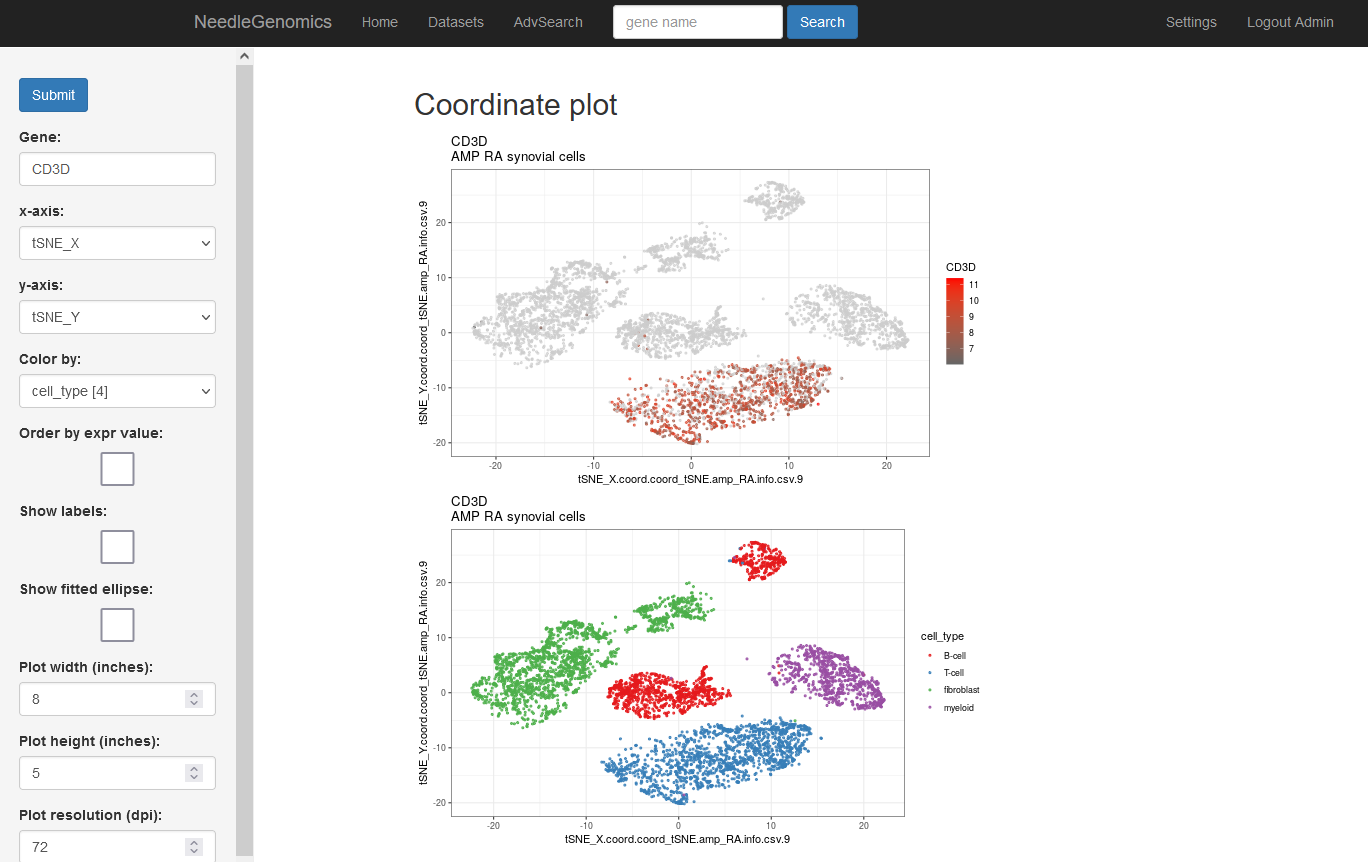
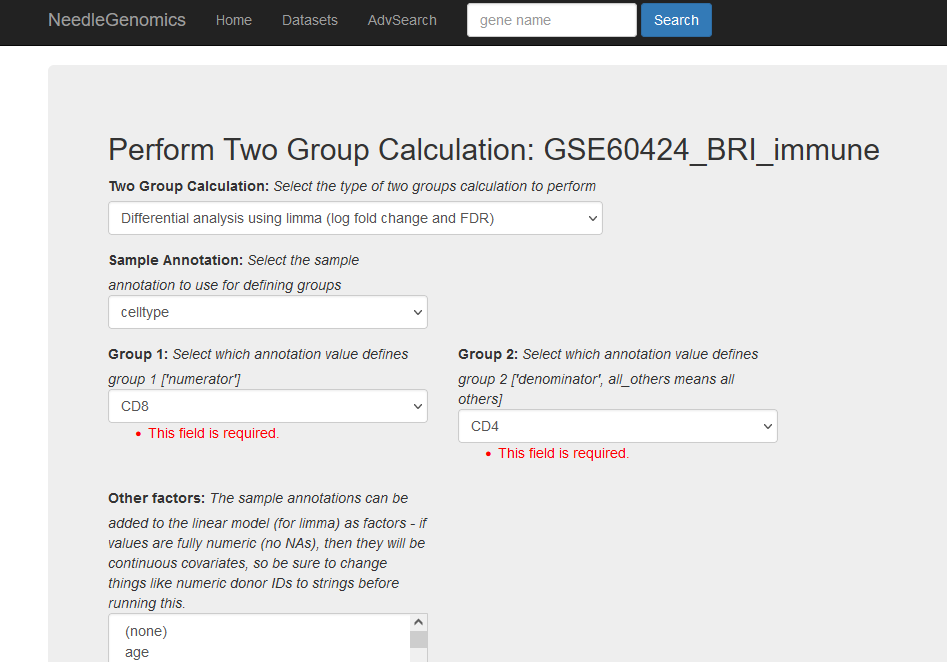
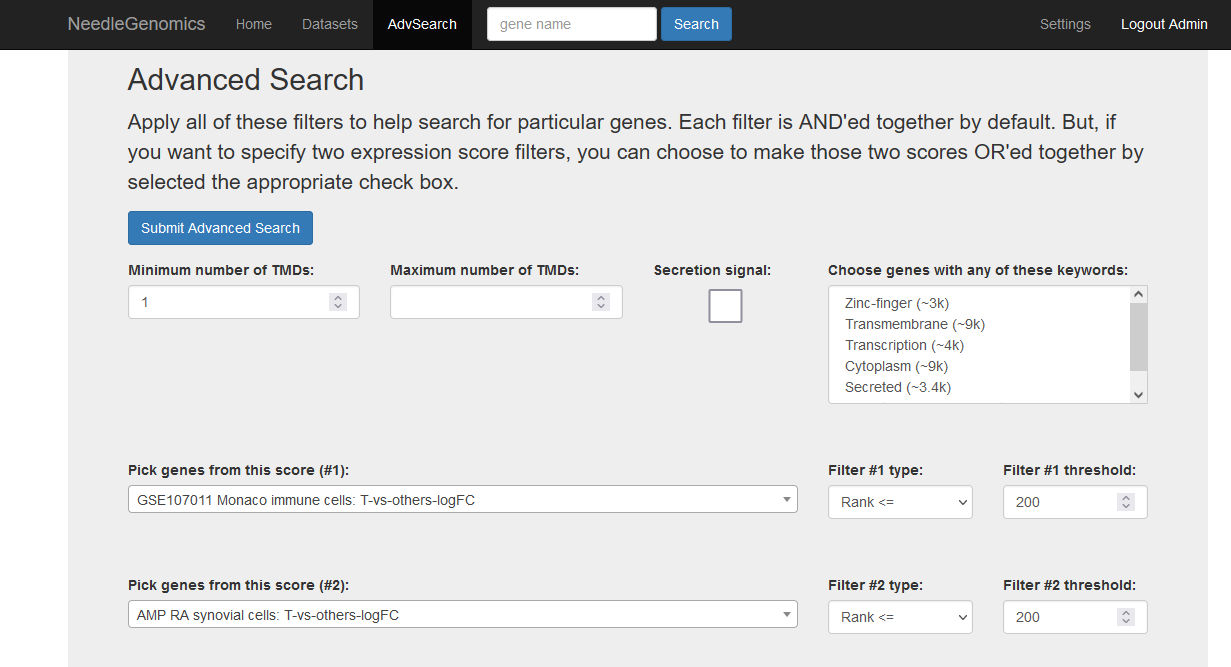
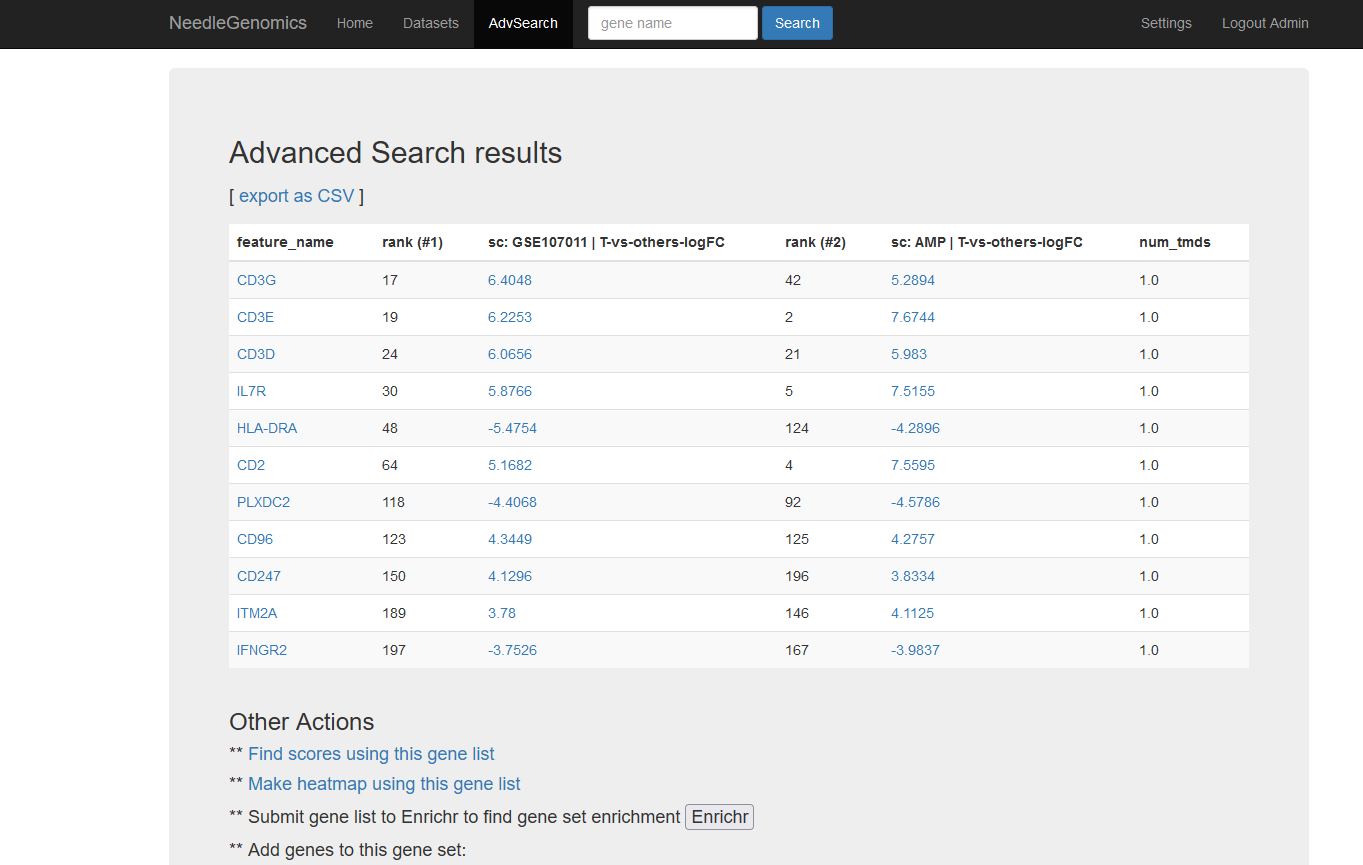
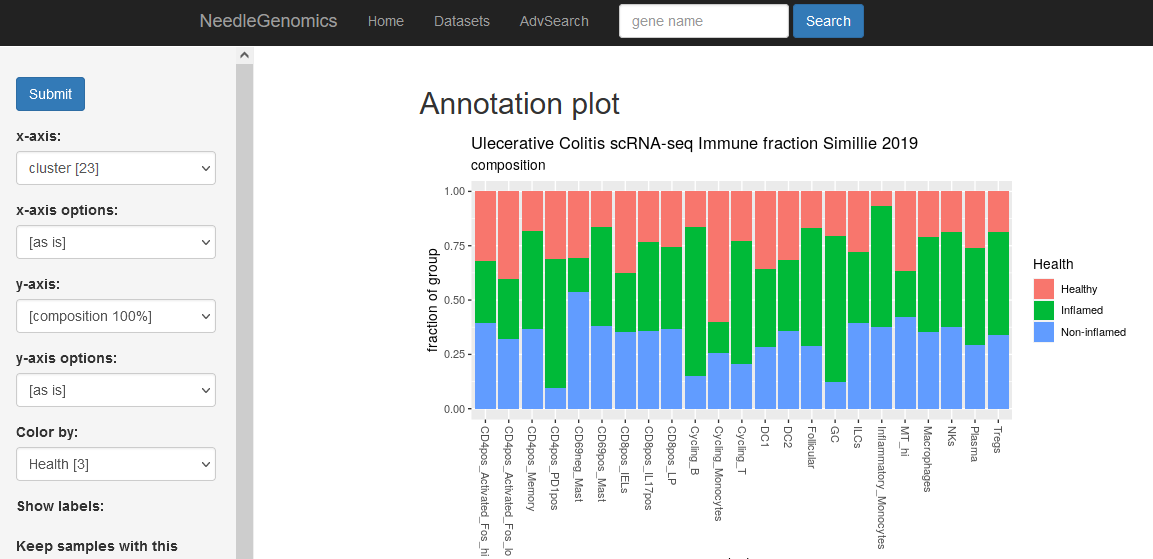
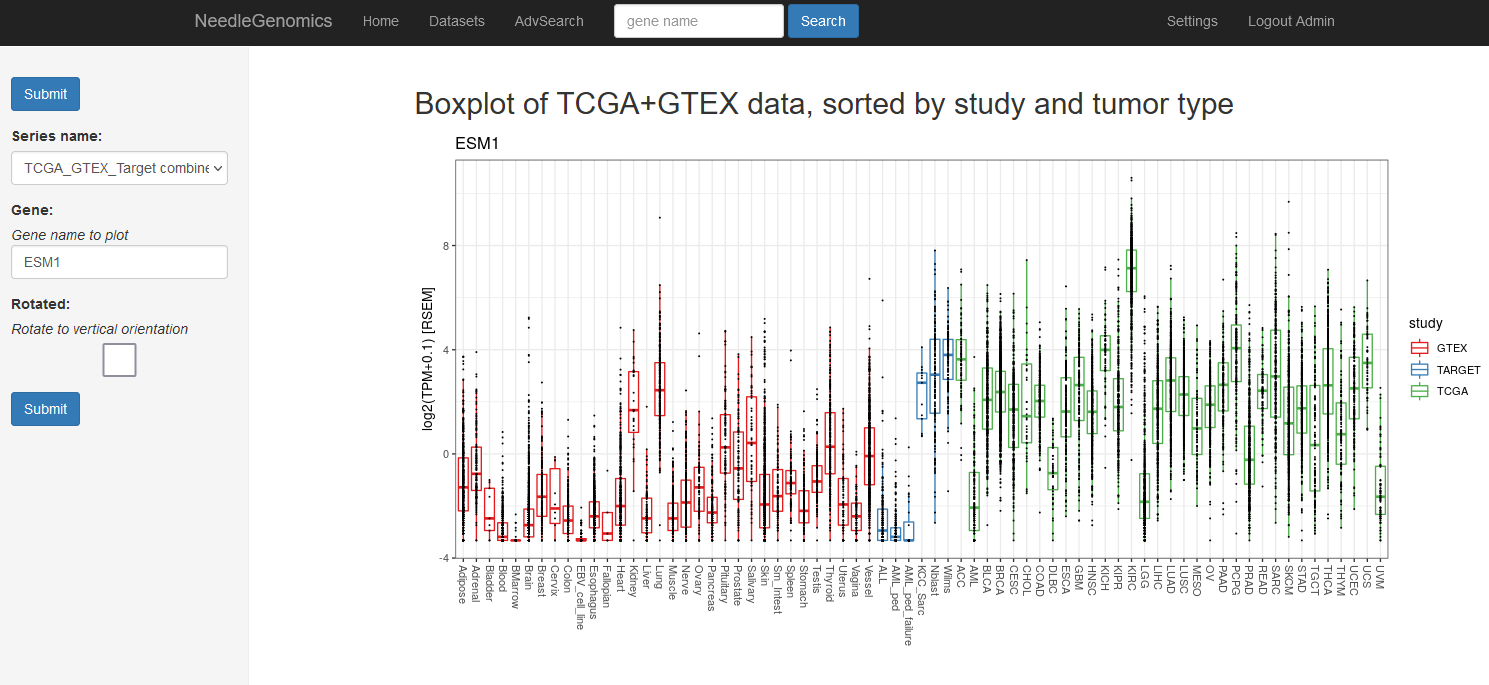
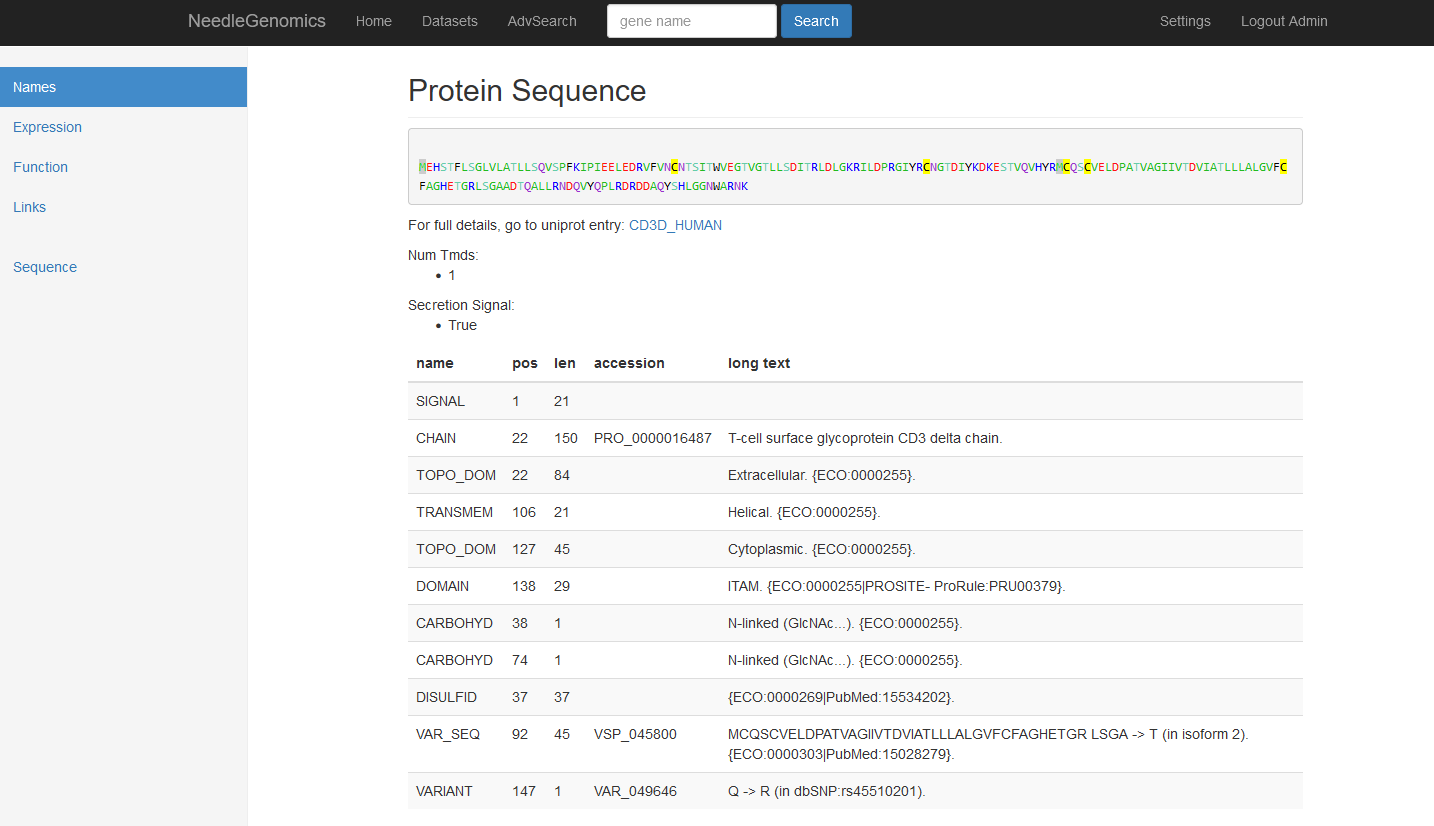
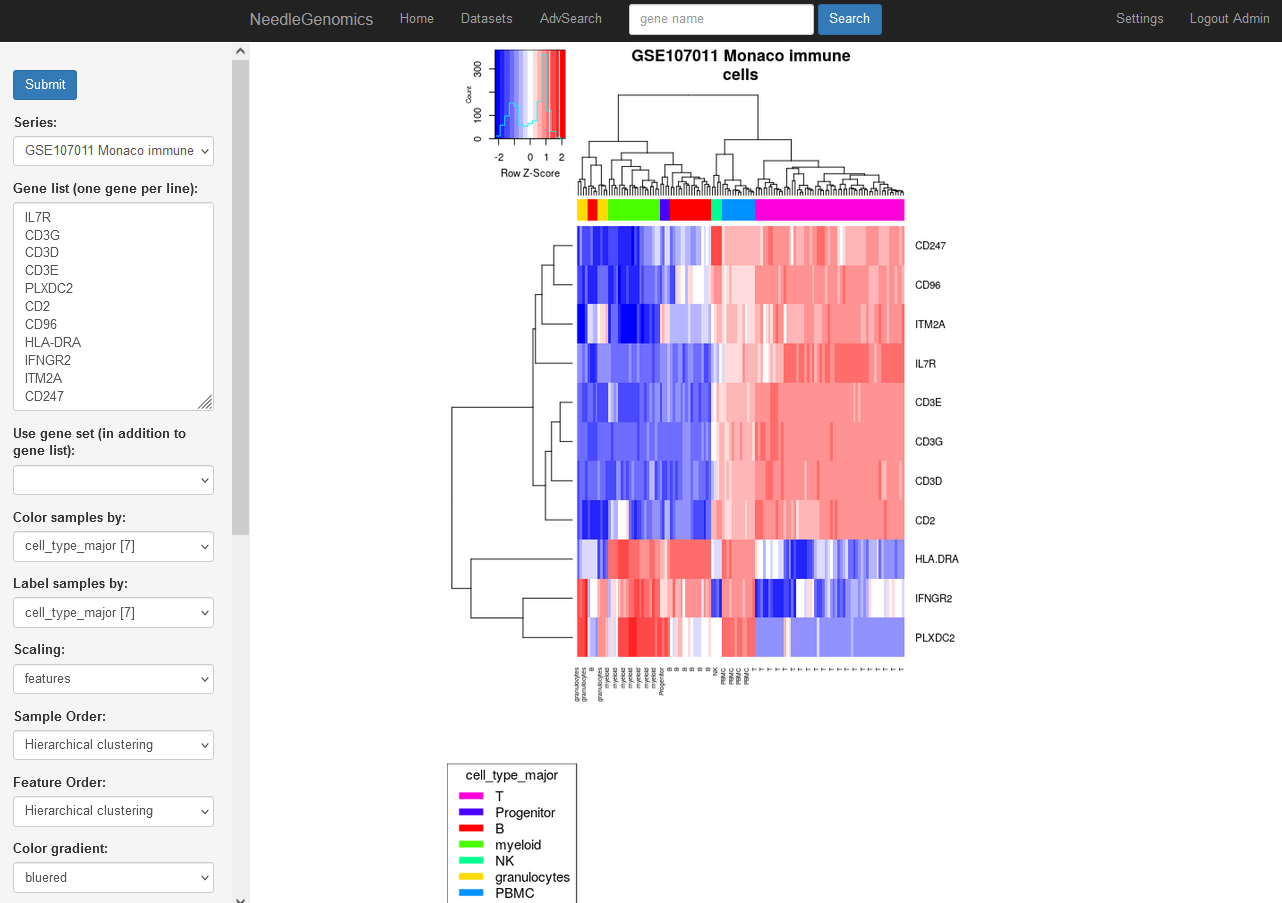
Screenshots
There are numerous ways to analyze and curate the datasets in the app.


Why needle?
- 01. Patients
- We really want you to succeed in making drugs or improving patients lives. This is our top priority
- 02. Discovery
- Target discovery is hard and a rare gift when it succeeds. We want to give more scientists the chance to find something new
- 03. Seattle
- We started a few miles from the iconic Needle in Seattle. We have global reach
Our Blog
The latest news in our blog
Contact
Reach out for a demo or licensing information. We won't spam you. We don't have pushy sales (we want to make sure your use case is a good fit)
Phone
[submit form first]
Home base
Seattle, WA
Email
brian at needlegenomics dot com